Software Solutions
ClusterEx:
Hit
lists consisting of genes or proteins are the output of many
genome-wide technologies. ClusterEx helps to cluster these hit lists
according with the help of coexpression database. We established a webserver version of this tool:
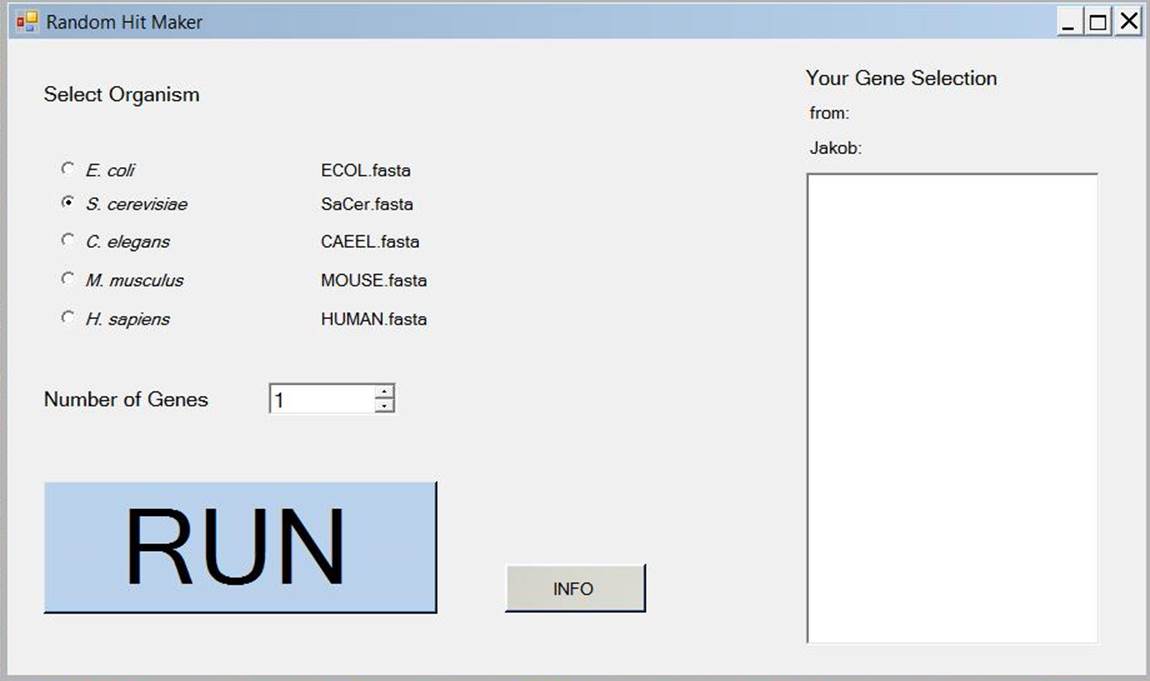
Random Hit Maker:
Methods like Microarrays, RNA sequencing or mass spectrometry yield lists of "hits", which can be further analysed to obtain interaction networks. In some cases control networks are needed, consisting of random gene lists to challenge the analysis algorithms. Random Hit Maker can be used to generate these random lists of genes for different organisms. It has been integrated into ClusterEx by now.
xMass
xMass helps with the analysis of peak lists from LC/MS Mass Spectrometric Data Sets in particular after crosslinking with isotope-labelled crosslinkers has been performed before.
diffUZ
dcdt-Plots can be used to quantitatively and qualitatively determine protein-protein interactions after analytical ultracentrifugation runs. diffUZ helps to streamline this process by generating ASCII-outputs on dc/dt plots for all samples in one rotor set.